Molecular Glue Degraders: Origins, Mechanisms, and Discovery Strategies
Get a free topical map and start building content authority today.
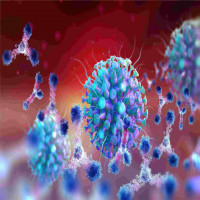
The term molecular glue degraders describes a class of small molecules that promote the proximity of a target protein to an E3 ubiquitin ligase, triggering ubiquitination and proteasomal degradation. Interest in molecular glue degraders has grown after discoveries linking immunomodulatory drugs to targeted degradation and the emergence of targeted protein degradation as a drug-discovery paradigm.
- Molecular glue degraders are small molecules that induce a neomorphic interaction between a target protein and an E3 ubiquitin ligase.
- Initial discoveries were largely serendipitous and followed by structure-guided studies that clarified mechanisms.
- Discovery approaches include phenotypic screening, chemoproteomics, structural biology and computational design.
- Challenges include selectivity, identification of compatible E3 ligases, resistance mechanisms and safety profiling.
History and early evidence
Serendipitous beginnings
Early evidence for small molecules that alter E3 ligase substrate specificity emerged from studies of immunomodulatory drugs in the late 20th and early 21st centuries. Compounds in that chemical class were found to bind cereblon (CRBN), a substrate receptor of the CUL4-RBX1-DDB1 E3 ligase complex, and to promote degradation of specific transcription factors. These observations framed a new concept: small molecules can act as adaptors to reprogram E3 ligases to degrade proteins that are not native substrates.
From observation to mechanism
Structural biology and biochemical studies showed that some small molecules create a composite surface that simultaneously engages an E3 ligase and a protein of interest, stabilizing a ternary complex and enabling ubiquitin transfer. This mechanism distinguishes molecular glue degraders from bifunctional degraders such as PROTACs, which connect ligase and target with a linker rather than creating a new interface.
Molecular glue degraders: mechanism and biology
How molecular glue degraders work
Mechanistically, a molecular glue degrader binds to an E3 ligase or a regulatory subunit and either directly recruits a new substrate or stabilizes a weak, transient interaction between ligase and substrate. Once the target protein is brought into proximity with the E3 catalytic core, ubiquitin is appended through the ubiquitin-proteasome system, marking the protein for degradation by the 26S proteasome. Key molecular players include the E1-E2-E3 cascade, substrate receptors (for example, CRBN or other substrate adaptors), and the proteasome.
Biological consequences
Degradation of a disease-relevant protein can lead to effects different from simple inhibition: complete removal of protein function, potential modulation of scaffolding roles, and altered signaling dynamics. These distinctive outcomes have motivated research into oncology, inflammation, and other therapeutic areas. Research programs also examine the cellular proteome-wide consequences of induced degradation, using proteomics and ubiquitinomics to profile on- and off-target effects.
Strategies used in the discovery of molecular glue degraders
Phenotypic screening and cell-based assays
Many molecular glue degraders were first identified through phenotypic screens that measure cell growth, differentiation, or other functional readouts. Hits from such screens can then be interrogated using chemoproteomic methods to identify protein targets and affected pathways.
Chemoproteomics and target identification
Mass spectrometry-based proteomics, including thermal proteome profiling and affinity enrichment, helps map small-molecule interactions across the proteome. Proteome-wide ubiquitination profiling can reveal proteins whose ubiquitination increases upon compound treatment, suggesting degradation pathways.
Structural biology and rational design
X-ray crystallography and cryo-electron microscopy have been central in understanding ternary complex formation and guiding structure-based modification to improve potency and selectivity. Computational docking and molecular dynamics simulation also support design efforts, especially when aiming to create or stabilize a novel interface between ligase and substrate.
High-throughput and fragment-based approaches
High-throughput screening of focused libraries, fragment-based lead discovery, and DNA-encoded libraries contribute diverse small molecules with potential glue-like activity. Screening strategies often incorporate counterscreens to detect non-specific proteasome activation or general cytotoxicity.
Applications, challenges, and regulatory considerations
Therapeutic potential and current research directions
Research into molecular glue degraders targets proteins considered "undruggable" by traditional modalities, including transcription factors and scaffolding proteins. Ongoing preclinical and clinical studies evaluate safety, dosing, and long-term effects of targeted degradation strategies. Regulatory agencies such as the U.S. Food and Drug Administration (FDA) and the European Medicines Agency (EMA) oversee clinical development and require robust characterization of mechanism, specificity, and toxicology.
Scientific and practical challenges
Key challenges include identifying suitable E3 ligases for particular targets, predicting or avoiding off-target degradation, overcoming acquired resistance (for example, via mutation or downregulation of the recruited E3 ligase), and ensuring favorable pharmacokinetics. Safety assessment must include proteome-wide monitoring and evaluation of secondary effects related to protein loss in tissues.
Research ethics and data transparency
Open sharing of structural data, proteomic findings and negative results accelerates understanding of which chemotypes and ligase combinations produce selective degradation. Academic institutions, funding agencies, and journals commonly encourage data deposition in repositories and adherence to reproducibility standards.
Tools and resources for researchers
Key experimental methods
Useful techniques include ubiquitin remnant profiling, tandem mass tag (TMT) quantitative proteomics, thermal shift assays, cellular thermal shift assay (CETSA), and cryo-EM for visualizing ternary complexes. Genetic tools such as CRISPR screens can help define cellular dependencies and resistance mechanisms.
Authoritative literature and databases
For comprehensive reviews and primary research papers on targeted protein degradation and molecular glue degraders, consult peer-reviewed literature and public biomedical databases. Selected reviews compile evidence on mechanisms, discovery methods and clinical translation; one accessible review on targeted protein degradation is available via PubMed Central for further reading: Targeted protein degradation: expanding the toolbox.
Outlook
Advances in chemical biology, structural determination and proteomics are likely to expand the repertoire of molecular glue degraders. Continued methodological innovation and careful evaluation of safety and selectivity will determine how broadly this approach can be applied across therapeutic areas.
What are molecular glue degraders?
Molecular glue degraders are small molecules that facilitate an interaction between a target protein and an E3 ubiquitin ligase, leading to ubiquitination and proteasomal degradation of the target.
How were molecular glue degraders discovered?
Discovery combined serendipitous clinical observations, phenotypic screening, structural biology, and chemoproteomics. Studies of immunomodulatory drugs that alter cereblon substrate specificity provided a concrete example that spurred focused discovery efforts.
How do researchers identify new molecular glue candidates?
Researchers use phenotypic screens, chemoproteomic target identification, structural studies, fragment-based methods and computational modeling, supported by proteomics to confirm target degradation.
What are the main safety concerns?
Safety concerns include off-target degradation, unintended loss of protein function in healthy tissues, and potential adaptive resistance. Regulatory assessment and proteome-wide profiling are important components of preclinical evaluation.
Where to find reliable scientific information?
Reliable information is available in peer-reviewed journals, public databases such as PubMed and PubMed Central, and guidance documents from regulatory agencies including the FDA and EMA.